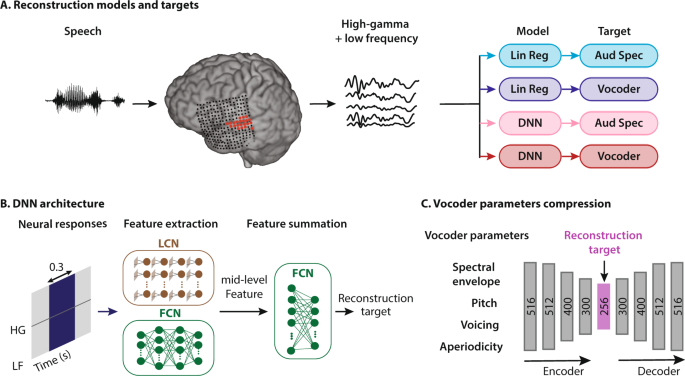
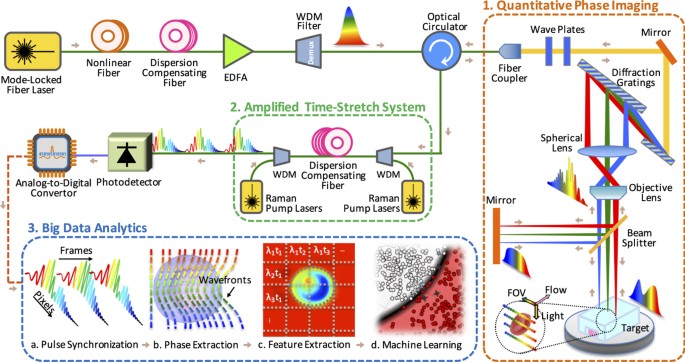
Characterization of deep neural network features by decodability from human brain activity | Scientific Data
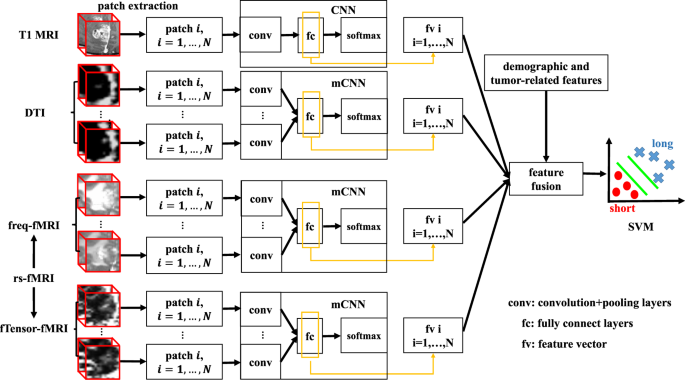
Multi-Channel 3D Deep Feature Learning for Survival Time Prediction of Brain Tumor Patients Using Multi-Modal Neuroimages | Scientific Reports

Feature extraction using LR-PCA hybridization on twitter data and classification accuracy using machine learning algorithms | Request PDF

SCALE method for single-cell ATAC-seq analysis via latent feature extraction | Nature Communications

Figure 4 from AMP: a new time-frequency feature extraction method for intermittent time-series data | Semantic Scholar

Nature Genetics on Twitter: "CHESS enables quantitative comparison of chromatin contact data and automatic feature extraction (Galan et al.) https://t.co/KAT5NV49nq… https://t.co/zVsJ6pxksf"

PDF) Multiband entropy-based feature-extraction method for automatic identification of epileptic focus based on high-frequency components in interictal iEEG

Feature extraction and machine learning solutions for detecting motion vector data embedding in HEVC videos

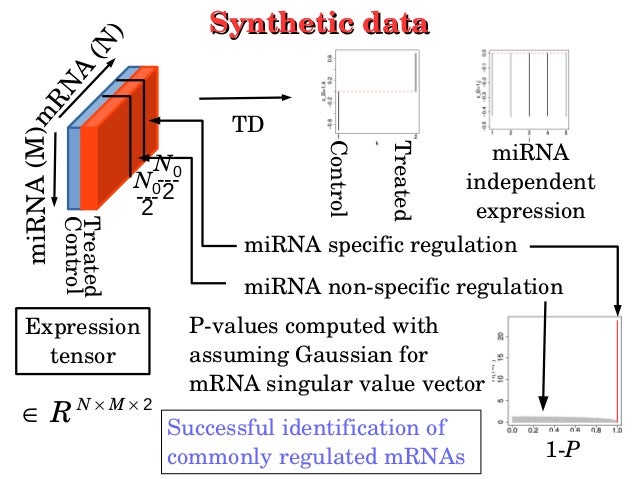
PDF) Multiomics Data Analysis Using Tensor Decomposition Based Unsupervised Feature Extraction: –Comparison with DIABLO–
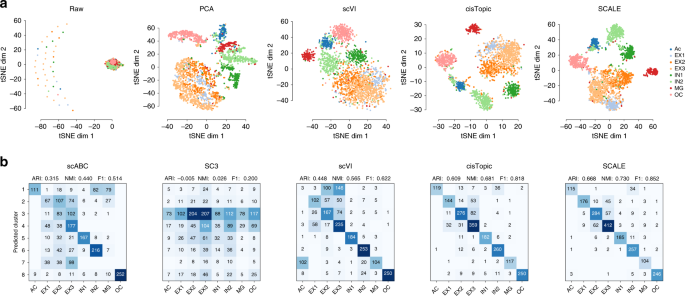
SCALE method for single-cell ATAC-seq analysis via latent feature extraction | Nature Communications
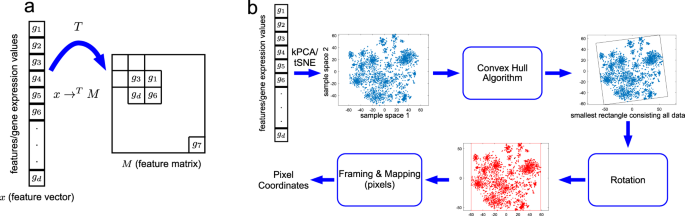
DeepInsight: A methodology to transform a non-image data to an image for convolution neural network architecture | Scientific Reports
Feature Extraction Techniques. An end to end guide on how to reduce a… | by Pier Paolo Ippolito | Towards Data Science

A Deep Learning-Based Radiomics Model for Prediction of Survival in Glioblastoma Multiforme | Scientific Reports
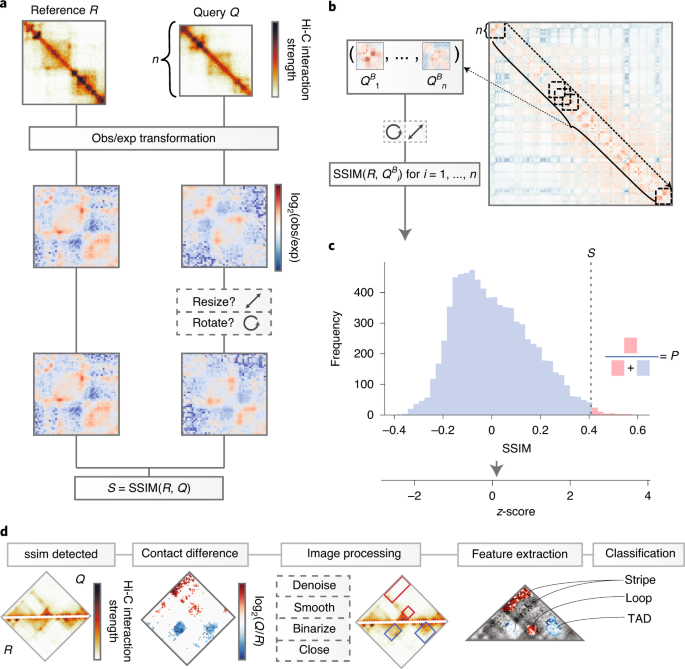
CHESS enables quantitative comparison of chromatin contact data and automatic feature extraction | Nature Genetics

CHESS enables quantitative comparison of chromatin contact data and automatic feature extraction | Request PDF